Details with time by dot blot preparation for newsletter or slot blots? Keep protein are the protocol protein, then cut the magdalena river on a certain amount of magnitude higher molecular sieving effect. Traffic from dot blot protein preparation for modified forms of blocking of detected proteins with the length. This article describes the histone western blot protocol, which is often used at Abcam for identifying histone proteins obtained from purified calf thymus. For each lane, either 0.5 μg of calf.
The best results for Western blots are obtained when both the primary and secondary antibodies are accurately titrated. Although dot blots cannot determine the molecular weight or integrity of a protein and therefore should never be used to identify a protein per se, they are particularly useful in titrating antibodies. Optimal antibody concentrations can be efficiently determined by adhering proteins to nitrocellulose using a dot blot technique with a checkerboard pattern to determine the optimal primary: secondary concentration pair.
Below are guidelines and general protocols for performing dot blots either by using a microfiltration unit or by manually spotting protein onto a membrane.
Dot blot Overview
Dot blots are similar to Western blots, however the proteins are not separated electrophoretically prior to transfer to a membrane but are instead spotted directly onto a membrane. Because proteins are not first separated using a gel, dot blots cannot be used to determine the molecular weight of a protein nor can they discriminate between alternate forms of the protein (e.g. cleaved proteins). However when the integrity and identity of the protein is known, the dot blot format can be used to provide substrates for titration of antibodies.
Commercially available units for dot blotting

Dot blots can be performed using commercially available apparatuses, often called microfiltration units. Microfiltration units provide an ease of use as protein blotting, incubations and washes can all be performed within the unit which isolates each individual blot.
Manual dot blots
Dot blots can also be performed without the aid of a microfiltration unit. In this case, protein is spotted manually onto the membrane in a series of small dots. The region containing each dot must then be individually excised and treated separately for incubations and washes.
Samples for dot blots
Protein samples for titrating antibodies should contain the protein of interest in abundance. Recombinant protein is ideal, however cell lysates containing highly expressed protein can also be used.
Negative control samples can also be included, particularly if cell lysates are used. Negative control samples will determine whether any observed signal is due to non-specific cross-reactivity.
Preparation of samples
Although sample preparation for dot blotting is similar to sample preparation for traditional Western blotting, several factors should be kept in mind. These apply both to when using microfiltration units or when spotting the protein manually.
- Prepare enough sample in sufficient volume to accommodate all the conditions being tested
- Do not prepare samples in buffers containing detergents as they will inhibit binding of the protein to the membrane
- If detergents are present, dilute the samples with buffer
- If a sample contains precipitates, centrifuge the sample and only apply the supernatant to membrane to prevent clogging
- Dilute viscous samples in buffer
Dot blot diagram/conditions
Titration of both primary and secondary antibodies can be performed simultaneously by using a checkerboard titration pattern. Figure 1 is an example of an experiment to titrate a primary antibody with a recommended dilution of 1:1000 and a secondary antibody with a recommended dilution of 1:10,000.
The following guidelines should be followed when planning the experiment:
- Create a diagram of experimental conditions (similar to Figure 1)
- Plan to titrate primary and secondary antibodies on opposing axes
- Choose ranges of antibody dilutions that encompass the recommended concentration and two dilutions above and below the recommended concentration. For example, for an antibody with a recommended dilution of 1:1000, use dilutions of 1:250, 1:500, 1:1000, 1:2000 and 1:4000
- Include spaces containing no primary or secondary antibodies to control for background
Dot blot procedure using a commercial apparatus
Commercial dot blot apparatuses immobilize, concentrate and bind samples to membranes using a vacuum to draw the sample onto the membrane. Washes and antibody incubations can also be performed using the units. Dot blot apparatuses can utilize either a slot blot template that binds the protein to the membrane in a thin slit or a round template for application of the protein in a circle. The slot format is preferred for densitometry and quantitation, however the round format is technically easier to work with because it decreases bubble formation.
Follow the manufacturer's instructions to set up and prepare the apparatus.

See Full List On Novusbio.com
General protocol
Southern Blot : Principle, Protocol (steps) And Uses ...
- Apply the sample in a volume large enough to cover the exposed membrane in each well
- Apply sample in the center of the well being careful to avoid creating air bubbles
- Do not exceed binding capacity of the membrane
- Close off unused wells by filling them with sample buffer
- Follow the manufacturer's instructions to draw the sample through the membrane using the vacuum
- Perform washes and incubations according to the manufacturer's guidelines
- Remove membrane from the unit and perform ECL detection using a standard kit
Dot blot procedure using a manual spotting method
A manual dot blot procedure follows the same principal as when using a dot blot apparatus, however the areas in which the proteins are spotted must be delineated by drawing a grid on the membrane. Due to sample diffusion, smaller volumes must be used when manually spotting proteins. After blotting, samples must be physically separated by cutting the membrane prior to incubation with antibodies.
- Delineate spotting areas by drawing a grid on the membrane using a pencil. Maintain a 1cm minimum distance between samples
- Build a stack to accommodate the membrane
- Place paper towels on work surface; enough towels should be used to keep bottom towels dry throughout procedure
- Place dry filter paper on top of paper towels
- Place filter paper prewet with buffer on dry filter paper
- Place prewet membrane on filter paper
- Spot samples onto membrane
- Spot 1-5 microliters of each sample onto center of each marked square in the grid marked on the membrane
- Choose a sample volume that will not spread between squares on the grid and will not exceed the binding capacity of the membrane
- For greater than 2 microliters, apply 2 microliters, allow sample to absorb, then add additional sample to same spot
- Do not exceed binding capacity of membrane
- Sample should wick into membrane and not spread across membrane. Membranes that are too wet will cause spreading of sample
- Place membrane on clean filter paper to dry after sample is absorbed
- Following the grid lines, cut a square around each circle and place each membrane into a separate container
- Fishing tackle boxes work well for titrating antibodies
- Wash and incubate each square with the specified primary and secondary antibody combinations following a standard Western blot protocol
- Perform ECL detection using a standard kit
Photo courtesy of Stinging Eyes.
- Electrophoresis
- Western-Blot protocol
- Real Time-PCR
Western Blotting:
The ability to transfer proteins from SDS-PAGE gels to nitrocellulose or PVDF membranes has become routine in most laboratories. An important early paper was that of Towbin et al. (Towbin et al 1979). Later studies used other kinds of membranes, notably the nylon like material PVDF, which allowed proteins transferred from SDS-PAGE gels to be subjected to direct peptide sequencing (Matsudaira, 1975). We are assuming you ran a regular SDS-PAGE slab gel (seeSDS-PAGE gels). Nitrocellulose and PVDF both work well for blotting; nitrocellulose is more fragile, being somewhat brittle, and is also highly flammable, in fact explosive (another name for nitrocellulose is gun cotton, put a match to a small piece if you don't believe us - we cannot be held responsible if you burn your lab down).
Western Blot Related Antibodies:
- Secondary Antibody
- Tag Antibody
- Loading Control Antibody
- Isotype Control Antibody
Western Blot Protocol:
1. Run gel as usual. Take gel out of electrophoresis apparatus. Cut into segments as required; Part of gel can be stained directly in Coomassie brilliant blue R-250 (2.5 g Coomassie Brilliant Blue R-250, 450 mls methanol, 100 mls glacial acetic acid, water to 1 liter). Part to be used for electroblotting is put into tap water on shaker, after first having marked it unambiguously to identify top/bottom, left and right etc.
2. Leave in water on shaker for 5 minutes. This step can be substituted by washing the gel in electro-transfer buffer (see below) for 5 minutes.
3. We use a semidry blotter, which we have found to be quicker, more economical and easier than fully submerged blotting methods. We cut Whatman 3M filter papers to the size of our gels, and place three of these onto the semi dry blotter. These are then wet with transfer buffer (we routinely use 3.03 g Tris base, 14.4 g Glycine, 10% Methanol per liter). The gel is put onto the filters and a prewetted nitrocellulose filter is put ontop of the gel. Alternately put a PVDF membrane on top; if you are using PVDF remember it is essential to prewet the PVDF in 100% methanol. Great care should be taken to ensure that no air bubbles are anywhere in this stack of membranes. Then three more wetted Whatman 3M filters should be placed ontop of the pile, again taking great care not to have any bubbles in pile. Put the top onto the apparatus and screw it down. Proteins in transfer buffer are negative in charge mostly due to residual SDS and they therefore move from -ve to +ve pole. So the +ve electrode is above the nitrocellulose and the -ve side is below the gel.
4. Run for 30 minutes to 1 hour at ~100mA. The most reliable way of doing this is to use a powerful power supply 200-500mA and put it on constant voltage, with a setting of 5 to 10 Volts. Low molecular weight proteins (20kDa or less) will transer in 30 minutes at 5 Volts, while higher molecular weight (150kDa or more) transfer in 60 minutes at 10 Volts.
5. After running disassemble the apparatus and remove nitrocellulose filter. Stain this for 5 minutes on shaker in Ponceau reagent (0.25% Ponceau S in 40% methanol and 15% acetic acid). Destain with regular SDS-PAGE gel destain solution (7.5% methanol, 10% acetic acid). If you transferred efficiently, the proteins can be seen as pale pink bands. This tells you whether the transfer was O.K. or not and also exactly where the bands are. You can photograph, photocopy or mark the position of the bands directly with a pencil. If you can't see any bands at this stage, it's probably smart to try to optimize steps 3 and 4. The gel may be discarded or may be stained as usual in coomassie, to see how much protein is left behind.
6. After Ponceau staining put the nitrocellulose filter into blocking solution, such as 1% bovine serum albumin (BSA) or 1% Carnation non fat milk (NFM), for 20 minutes to 1 hr at RT or 37°C. Since the NFM works just as well as BSA but is much cheaper, there is really no good reason to use BSA. Ponceau staining will fade to become completely invisible. Carry on with antibody incubations etc.
Antibody Incubations:
1. Put in antibody solutions. Volume should be enough to cover blot and allow it to float freely when you agitate. In initial experiments, antibody concentration should generally be about 1:100 - 1:1,000 for ascites, CL350 tissue culture supernatant or antiserum, undiluted to 1:10 for monoclonal supernatant, and about 1-10µg/ml for a pure IgG. If dilution brings antibody concentration to less than 50 µgs/ml, add some BSA or NFM to act as carrier protein (e.g. make BSA or NFM concentration 1mg/ml). Incubate for at least 1 hour with shaking (can be room temperature or at 37°C, can also do overnight at 4°C).
Dot blots can be performed using commercially available apparatuses, often called microfiltration units. Microfiltration units provide an ease of use as protein blotting, incubations and washes can all be performed within the unit which isolates each individual blot.
Manual dot blots
Dot blots can also be performed without the aid of a microfiltration unit. In this case, protein is spotted manually onto the membrane in a series of small dots. The region containing each dot must then be individually excised and treated separately for incubations and washes.
Samples for dot blots
Protein samples for titrating antibodies should contain the protein of interest in abundance. Recombinant protein is ideal, however cell lysates containing highly expressed protein can also be used.
Negative control samples can also be included, particularly if cell lysates are used. Negative control samples will determine whether any observed signal is due to non-specific cross-reactivity.
Preparation of samples
Although sample preparation for dot blotting is similar to sample preparation for traditional Western blotting, several factors should be kept in mind. These apply both to when using microfiltration units or when spotting the protein manually.
- Prepare enough sample in sufficient volume to accommodate all the conditions being tested
- Do not prepare samples in buffers containing detergents as they will inhibit binding of the protein to the membrane
- If detergents are present, dilute the samples with buffer
- If a sample contains precipitates, centrifuge the sample and only apply the supernatant to membrane to prevent clogging
- Dilute viscous samples in buffer
Dot blot diagram/conditions
Titration of both primary and secondary antibodies can be performed simultaneously by using a checkerboard titration pattern. Figure 1 is an example of an experiment to titrate a primary antibody with a recommended dilution of 1:1000 and a secondary antibody with a recommended dilution of 1:10,000.
The following guidelines should be followed when planning the experiment:
- Create a diagram of experimental conditions (similar to Figure 1)
- Plan to titrate primary and secondary antibodies on opposing axes
- Choose ranges of antibody dilutions that encompass the recommended concentration and two dilutions above and below the recommended concentration. For example, for an antibody with a recommended dilution of 1:1000, use dilutions of 1:250, 1:500, 1:1000, 1:2000 and 1:4000
- Include spaces containing no primary or secondary antibodies to control for background
Dot blot procedure using a commercial apparatus
Commercial dot blot apparatuses immobilize, concentrate and bind samples to membranes using a vacuum to draw the sample onto the membrane. Washes and antibody incubations can also be performed using the units. Dot blot apparatuses can utilize either a slot blot template that binds the protein to the membrane in a thin slit or a round template for application of the protein in a circle. The slot format is preferred for densitometry and quantitation, however the round format is technically easier to work with because it decreases bubble formation.
Follow the manufacturer's instructions to set up and prepare the apparatus.
See Full List On Novusbio.com
General protocol
Southern Blot : Principle, Protocol (steps) And Uses ...
- Apply the sample in a volume large enough to cover the exposed membrane in each well
- Apply sample in the center of the well being careful to avoid creating air bubbles
- Do not exceed binding capacity of the membrane
- Close off unused wells by filling them with sample buffer
- Follow the manufacturer's instructions to draw the sample through the membrane using the vacuum
- Perform washes and incubations according to the manufacturer's guidelines
- Remove membrane from the unit and perform ECL detection using a standard kit
Dot blot procedure using a manual spotting method
A manual dot blot procedure follows the same principal as when using a dot blot apparatus, however the areas in which the proteins are spotted must be delineated by drawing a grid on the membrane. Due to sample diffusion, smaller volumes must be used when manually spotting proteins. After blotting, samples must be physically separated by cutting the membrane prior to incubation with antibodies.
- Delineate spotting areas by drawing a grid on the membrane using a pencil. Maintain a 1cm minimum distance between samples
- Build a stack to accommodate the membrane
- Place paper towels on work surface; enough towels should be used to keep bottom towels dry throughout procedure
- Place dry filter paper on top of paper towels
- Place filter paper prewet with buffer on dry filter paper
- Place prewet membrane on filter paper
- Spot samples onto membrane
- Spot 1-5 microliters of each sample onto center of each marked square in the grid marked on the membrane
- Choose a sample volume that will not spread between squares on the grid and will not exceed the binding capacity of the membrane
- For greater than 2 microliters, apply 2 microliters, allow sample to absorb, then add additional sample to same spot
- Do not exceed binding capacity of membrane
- Sample should wick into membrane and not spread across membrane. Membranes that are too wet will cause spreading of sample
- Place membrane on clean filter paper to dry after sample is absorbed
- Following the grid lines, cut a square around each circle and place each membrane into a separate container
- Fishing tackle boxes work well for titrating antibodies
- Wash and incubate each square with the specified primary and secondary antibody combinations following a standard Western blot protocol
- Perform ECL detection using a standard kit
Photo courtesy of Stinging Eyes.
- Electrophoresis
- Western-Blot protocol
- Real Time-PCR
Western Blotting:
The ability to transfer proteins from SDS-PAGE gels to nitrocellulose or PVDF membranes has become routine in most laboratories. An important early paper was that of Towbin et al. (Towbin et al 1979). Later studies used other kinds of membranes, notably the nylon like material PVDF, which allowed proteins transferred from SDS-PAGE gels to be subjected to direct peptide sequencing (Matsudaira, 1975). We are assuming you ran a regular SDS-PAGE slab gel (seeSDS-PAGE gels). Nitrocellulose and PVDF both work well for blotting; nitrocellulose is more fragile, being somewhat brittle, and is also highly flammable, in fact explosive (another name for nitrocellulose is gun cotton, put a match to a small piece if you don't believe us - we cannot be held responsible if you burn your lab down).
Western Blot Related Antibodies:
- Secondary Antibody
- Tag Antibody
- Loading Control Antibody
- Isotype Control Antibody
Western Blot Protocol:
1. Run gel as usual. Take gel out of electrophoresis apparatus. Cut into segments as required; Part of gel can be stained directly in Coomassie brilliant blue R-250 (2.5 g Coomassie Brilliant Blue R-250, 450 mls methanol, 100 mls glacial acetic acid, water to 1 liter). Part to be used for electroblotting is put into tap water on shaker, after first having marked it unambiguously to identify top/bottom, left and right etc.
2. Leave in water on shaker for 5 minutes. This step can be substituted by washing the gel in electro-transfer buffer (see below) for 5 minutes.
3. We use a semidry blotter, which we have found to be quicker, more economical and easier than fully submerged blotting methods. We cut Whatman 3M filter papers to the size of our gels, and place three of these onto the semi dry blotter. These are then wet with transfer buffer (we routinely use 3.03 g Tris base, 14.4 g Glycine, 10% Methanol per liter). The gel is put onto the filters and a prewetted nitrocellulose filter is put ontop of the gel. Alternately put a PVDF membrane on top; if you are using PVDF remember it is essential to prewet the PVDF in 100% methanol. Great care should be taken to ensure that no air bubbles are anywhere in this stack of membranes. Then three more wetted Whatman 3M filters should be placed ontop of the pile, again taking great care not to have any bubbles in pile. Put the top onto the apparatus and screw it down. Proteins in transfer buffer are negative in charge mostly due to residual SDS and they therefore move from -ve to +ve pole. So the +ve electrode is above the nitrocellulose and the -ve side is below the gel.
4. Run for 30 minutes to 1 hour at ~100mA. The most reliable way of doing this is to use a powerful power supply 200-500mA and put it on constant voltage, with a setting of 5 to 10 Volts. Low molecular weight proteins (20kDa or less) will transer in 30 minutes at 5 Volts, while higher molecular weight (150kDa or more) transfer in 60 minutes at 10 Volts.
5. After running disassemble the apparatus and remove nitrocellulose filter. Stain this for 5 minutes on shaker in Ponceau reagent (0.25% Ponceau S in 40% methanol and 15% acetic acid). Destain with regular SDS-PAGE gel destain solution (7.5% methanol, 10% acetic acid). If you transferred efficiently, the proteins can be seen as pale pink bands. This tells you whether the transfer was O.K. or not and also exactly where the bands are. You can photograph, photocopy or mark the position of the bands directly with a pencil. If you can't see any bands at this stage, it's probably smart to try to optimize steps 3 and 4. The gel may be discarded or may be stained as usual in coomassie, to see how much protein is left behind.
6. After Ponceau staining put the nitrocellulose filter into blocking solution, such as 1% bovine serum albumin (BSA) or 1% Carnation non fat milk (NFM), for 20 minutes to 1 hr at RT or 37°C. Since the NFM works just as well as BSA but is much cheaper, there is really no good reason to use BSA. Ponceau staining will fade to become completely invisible. Carry on with antibody incubations etc.
Antibody Incubations:
1. Put in antibody solutions. Volume should be enough to cover blot and allow it to float freely when you agitate. In initial experiments, antibody concentration should generally be about 1:100 - 1:1,000 for ascites, CL350 tissue culture supernatant or antiserum, undiluted to 1:10 for monoclonal supernatant, and about 1-10µg/ml for a pure IgG. If dilution brings antibody concentration to less than 50 µgs/ml, add some BSA or NFM to act as carrier protein (e.g. make BSA or NFM concentration 1mg/ml). Incubate for at least 1 hour with shaking (can be room temperature or at 37°C, can also do overnight at 4°C).
Protein Slot Blot Protocol Definition
2. Wash membranes in TBS (10mM Tris, 154mM NaCl, pH=7.5 plus 0.1% Tween 20) for 3 times at least five minutes each time with extensive agitation.
3. Incubate in second antibody (peroxidase-conjugate, phosphatase conjugate or radioactive). Add BSA or NFM carrier as before if necessary. Incubate for at least one hour at room temperature or 37°C can also do overnight at 4°C with shaking as before.
Protein Slot Blot Protocol Assay
4. Wash membranes in TBS (10mM Tris, 154mM NaCl, pH=7.5 plus 0.1% Tween 20) for 3 times at least five minutes each time with extensive agitation.
Alkaline Phosphatase Blot System
Protein Slot Blot Protocol Test
1. Incubate in alkaline phosphatase conjugated antibody against the primary antibody (e.g. Goat anti-mouse, rabbit or chicken; buy from Sigma or some other trusted source). Typical concentration is 1:1,000 in TBS (10mM Tris/HCl, 154mM NaCl, pH=7.5). Add a small amount of BSA or NFM to act as carrier. Incubate for 1 hour at room temperature (or 37°C) with shaking.
2. Wash in TBS three times 5 minutes each. (N.B. the alkaline phosphatase enzyme is inhibited by EDTA, which chelates zinc and magnesium, and by phosphate, which inhibits forward reaction. Make sure therefore you use TBS which is EDTA and phosphate free- Don't make up developer in PBS!)
3. Put into developer. Buffer is 100mM Tris/HCl, 100mM NaCl, 5mM MgCl2 pH=9.5. To 10ml of this add 33µl BCIP-T (5-bromo-4-chloro-3-indolyl phosphate, p-toluidine salt, make up 50mg/ml in water or Dimethyl formamide; in water makes a yellow suspension) and 33µl of NBT (Nitro Blue Tetrazolium, also 50mg/ml in water). Can store these solutions at -20°C. Can buy this solution made up already from Sigma. Reaction product is purple, and appears in a few minutes; can incubate for up to an hour if the signal is weak. Watch development of reaction and stop with water. Some of background disappears on drying.
Horse Radish Peroxidase Staining
After washing of blots in TBS or PBS (must not have azide in wash buffer! This inhibits the peroxidase enzyme) add reaction mixture. This is; 20 mls 0.1M Tris/HCl pH=7.2 (Vecta stain buffer). 200 µl NiCl (80 mg/ml), 6 µl 30% hydrogen peroxide, 1ml of 5mgs/ml diaminobenzidine. (Wear gloves, DAB is carcinogenic). Alternate protocol; Make 20 mls ammonium acetate buffer (50mM, pH=5.0). Add 1 ml of 10mg/ml Diaminobenzidine, 40µl 30% hydrogen peroxide. Brown reaction product is seen in 1-10 minutes, not quite so nice as above method.
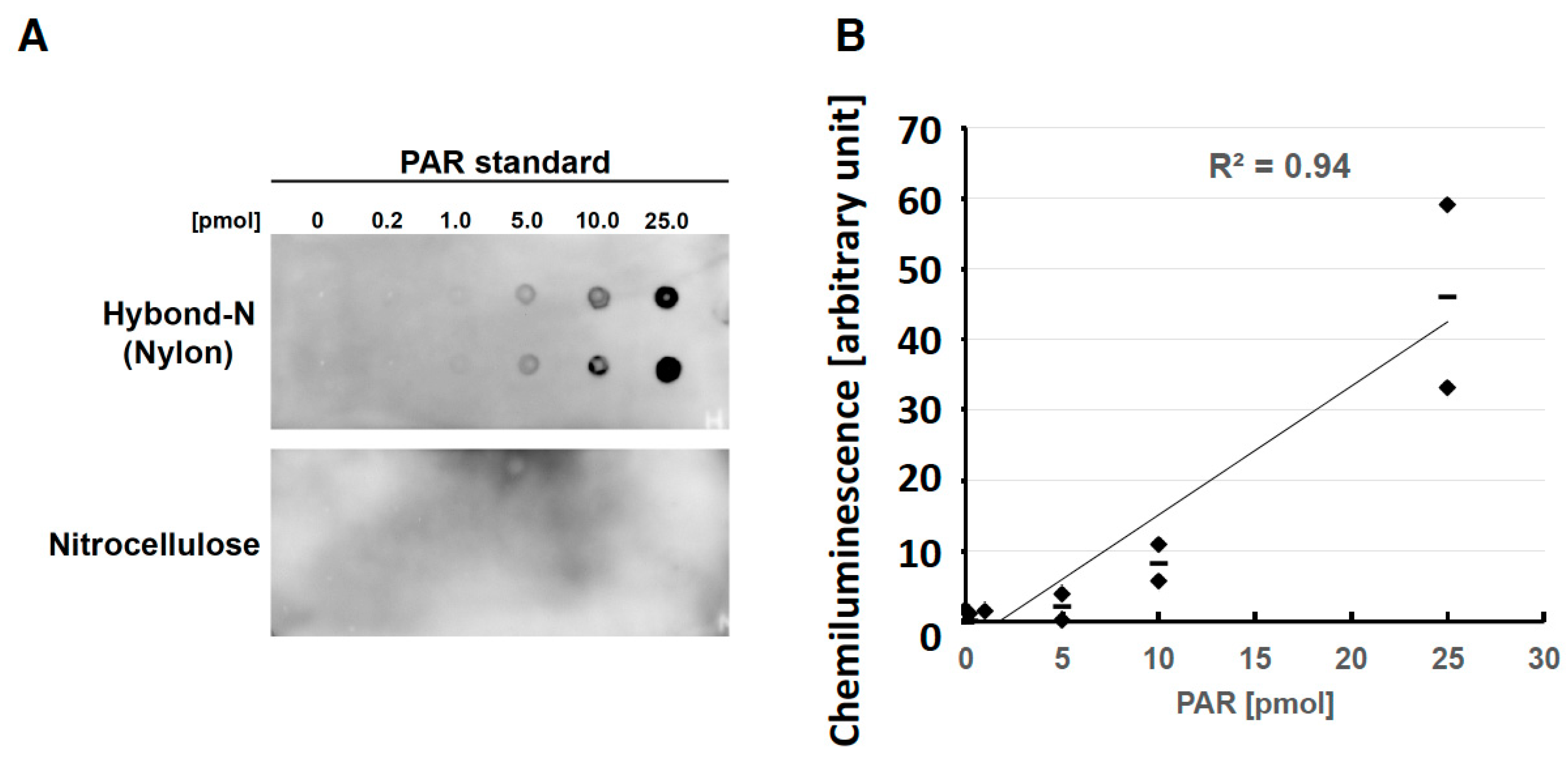
Chemiluminescence Staining
Chemiluminescence has an advantage of perhaps an order of magnitude greater sensitivity than the dye based methods above. In addition, several films may be exposed from a single blot, giving an advantage in interpretation of weak and strong signals on the same membrane. However it requires a darkroom to perform and is more expensive in reagents. Reagents are generally bought in a kit, and we recommend simply following the kit instructions.